Introduction
Peter Damerow focused on big data projects, digital collections, and computational tools long before digital humanities and big data became popular concepts. His pioneering efforts in these nascent fields were a logical consequence of his driving research questions: How can we understand the longue durée patterns in the history of knowledge? How can we understand the transitions from nomadic hunters and gatherers to early civilizations and what role did organized and abstract knowledge play in these transitions? What was the origin of writing and mathematics? What parts of the cognitive make-up of humans and early societies facilitated these transitions?
Clearly these are “big questions.” Answering them is by necessity a collective endeavor. And, as everybody who has organized an even remotely similar research project knows, sharing information, data, interpretations, being able to work collaboratively, and being able to connect evidence from different fields is absolutely crucial. But this does not happen by itself. It was Peter’s early vision to build the necessary infrastructure—from the earliest digital databases in the history of science to the most advanced open access publication platforms in our field.
In addition to sharing data and scholarly interpretations, Peter also had an active interest in and promoted the development of computational tools. For him, as a historian of early writing and mathematics, the possibilities of new algorithms that could analyze patterns of historical change or of new graph-based representations of knowledge were further steps in a process of knowledge acquisition that began deep in our evolutionary past. It is with a sense of deep gratitude and appreciation that we present a brief overview of some of the computational tools that we developed in order to analyze the complex patterns and processes within the history of knowledge. We are, in a fitting case of historical continuity, Peter’s daughter Julia, a computer scientist and, as of this summer (2014), a freshly minted PhD in computational history and philosophy of science; Erick Peirson, Julia’s congenial partner in the development of the computational research system introduced here, and a biologist and historian of science; and MDL, who is as proud of his academic children as Peter was of his daughter.
In Lunenfeld (2012), the authors observe that in digital humanities “[...] what we are seeing is the emergence of new conjunctions between the macro and the micro, general surface trends and deep hermeneutic inquiry, the global view from above and the local view on the ground” (Lunenfeld et al. 2012, 39). In contrast to close reading and careful studying of individual sources (the micro-scale), which are key methods in the humanities, distant reading1 in digital humanities employs computational methods to analyze large text corpora in order to find overall patterns, trends, or connections (the macro-scale) (Lunenfeld et al. 2012). Lunenfeld et al. see “zooming in and out” between distant and close reading as a powerful tool of digital humanists. Müller calls this process “scalable reading,” comparing it to the zoom function in Google Earth (Müller 2012). He states that scalable reading enables scholars to easily switch between the details of a text and its context (Müller 2012). Computers can support researchers by making vast amounts of data such as texts or images accessible through automatic extraction, analysis, and visualization of information. They can provide scholars with new tools that might help discover unknown relationships or patterns. However, they cannot replace the careful interpretation and examination of individual sources by a scholar.
This paper describes a research system called “Quadriga System” that is based on the idea of representing texts as networks of concepts that can be mathematically analyzed and visualized. These networks are created by scholars through close reading and structured annotation of texts. However, the Quadriga System follows a collaborative approach that facilitates the creation of a large-scale data repository to enable data-driven research in the history and philosophy of science. The system can therefore be placed in between the micro- and the macro-level of source analysis, on the so-called meso-level (or meso-scale). It is designed to help researchers detecting patterns and relationships of interest in their sources by transforming the materials into structured datasets on the micro-level and analyzing them on the macro-level. The Quadriga System follows a similar approach to projects in the field of bioinformatics such as GenBank that rely on different contributors from around the world to submit new entries to the database (Benson et al. 2010). The data structure underlying the Quadriga System (called Quadruples) enables scholars to seamlessly switch back and forth between a single text and a whole corpus, facilitating scalable reading.
In this paper, we will first briefly describe two projects that use the Quadriga System: the Genecology Project and the EP Annotation Project. We will then detail the system’s architecture and its different components. The last section will discuss how the history of science might benefit from using the Quadriga System.
The Genecology Project
The Genecology Project2 studies genecology research in Great Britain during the twentieth century. Genecology is a branch of ecology that studies how genetic differences in plant populations relate to “geospatial variation in environmental factors (e.g. soils, altitude, climate)” (Peirson, Damerow, and Laubichler forthcoming, 3). The project analyzes how the conceptual change that occurred in genecology research was influenced by contributing researchers and their interactions and collaborations with a focus on one particular researcher: Tony Bradshaw. It also asks how ideas and theories in the field spread, and how they changed. In its first phase, the Genecology Project is therefore especially interested in identifying the main actors contributing to genecology research and who collaborated with Bradshaw, and how the patterns of collaboration among the researchers changed over time. To answer this question the project concentrates on constructing a social network from interactions, collaborations, and the institutional contexts of genecology researchers.
The Genecology Project follows a text-driven approach that is not simply based on biographical information, but also relies on acknowledgment sections of publications or other textual evidence demonstrating collaborative efforts, such as co-authorship. Texts were selected based on an initial list of papers published in 1964 that provides an overview of genecology research at that time. In the first stage, all papers from that list were digitized and annotated. In a second step, publications cited by the listed papers or other manuscripts by listed authors and co-authors were analyzed as well.
The selected texts were annotated with a set of predefined relationships using a software application called Vogon. Vogon allows a researcher to create a certain kind of annotation that points to the position of a word in a text and a so-called “concept” that specifies what a word refers to. Those annotations can then be put in relation to one another. For example, if a text states that a person helped the author with a certain task, the author of the text as well as the person helping him will be annotated with concepts representing the two people. The two resulting annotations will then be connected by an “engages with” relationship or any other well-defined relationship the annotator choses. Similarly, the relationship between a researcher and his affiliated institution is expressed by an “employs” relation between two annotations representing the institution and the researcher. Several annotations of this format create a network of “concepts,” which in the case of the Genecology Project is a social network of persons and institutions.
Such a network can be exported from Vogon in a standard graph format such as XGMML to be visualized in a network visualization application (such as Cytoscape),3 or if geographical information is attached to the nodes of a network, it can be plotted on a map (see fig. 21.1). A visualization as shown in figure 21.1 facilitates quick processing of the displayed data by a viewer and can reveal information that otherwise might stay hidden (Mazza 2009).
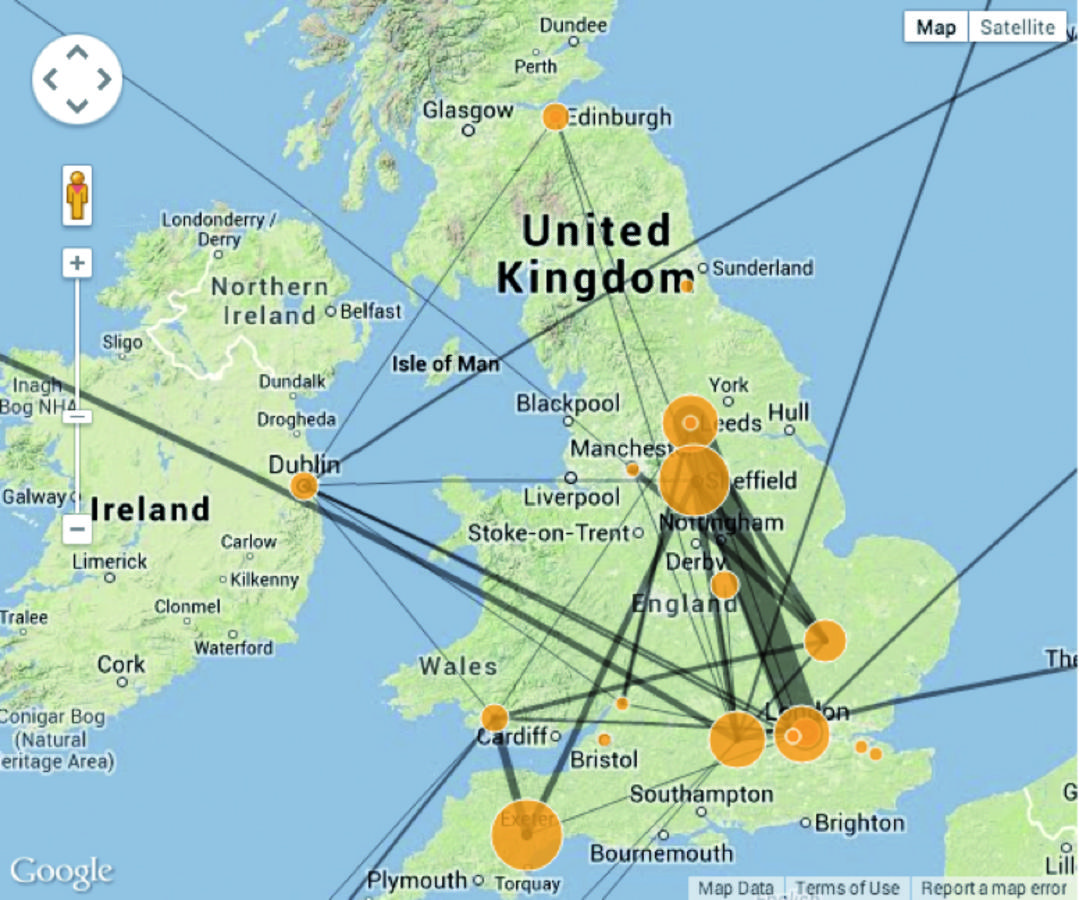
Fig. 21.1: Social Network created by the Genecology Project plotted on a map.
The EP Annotation Project
The goal of the EP Annotation Project is to annotate articles written for the Embryo Project with relationships that reflect how the entities described in the articles relate to each other. The Embryo Project is an online encyclopedia of embryology that aims to document embryo research in the broadest possible way (Laubichler and Maienschein 2009). Articles in the Embryo Projects “are written and marked up in such a way that they help populating the database with additional objects that have interesting and relevant relationships to the object of the entry” (Laubichler and Maienschein 2009, 11). For example, there exists an article about Hans Spemann that mentions that Spemann worked with Theodor Boveri and Wilhelm Röntgen. In the marked-up article this information is turned into annotations that represent “worked with” relationships between Spemann and Boveri and Spemann and Röntgen. However, while an entry for Boveri exists, there is no article about Röntgen. By creating relationships between Röntgen and other entities such as Spemann, information about Röntgen is stored and available for use although no article has been written yet. One motivation for creating and storing such relationships is to be able to easily answer questions such as “Who was a student of whom?” or “Who worked at a particular place? With what particular organisms?” (Laubichler and Maienschein 2009, 9).
As the EP Annotation Project is an exploratory project, it so far has been undertaken as a proof of concept project. There are about 50 articles that were annotated using Vogon; all of them describe specific persons (e.g., Hans Spemann or Viktor Hamburger) rather than institutions or organisms. For each article about 10 to 20 relationships were created, capturing information such as who was a teacher of whom, who worked with what organism, or what kind of relationship existed between a person and institution.
For the visualization of the annotations created with Vogon, annotations were transformed into graphs in which every node represents a concept of interest (for example a person or organism), and edges represent relationships between those concepts (i.e., “contributed to” or “used”). Figure 21.2 shows a network of people and organisms, techniques, or theories those people worked on. Such a network could be used to explore the articles in the Embryo Project by browsing through the concepts (represented by nodes) and their relationships to each other (edges). As Vogon allows text positions to be stored with annotations, a person using the network could jump directly to the texts that mention a specific relationship between, for example, a person and an organism. Moreover, when time information is added to the annotations, the networks resulting from the annotations could allow a user of the Embryo Project Encyclopedia to explore its content filtered by time or place, or to create timelines to visualize, for instance, who worked on a concept or theory over time.
The Quadriga System
The Quadriga System is based on the idea to represent texts as graphs. By representing unstructured texts as graphs, the information contained in a text is given a mathematical structure that can be used for computational analysis. The basic components of these graphs are so-called “Quadruples,” (see fig. 21.3) also referred to as “contextualized triples.” The basic idea is similar to a concept proposed in Macgregor and Ko (2003). Macgregor and Ko describe quads (a four-tuple consisting of subject, predicate, object, and context) in which a context can itself be part of a set of assertions that define the “environment” of that context.
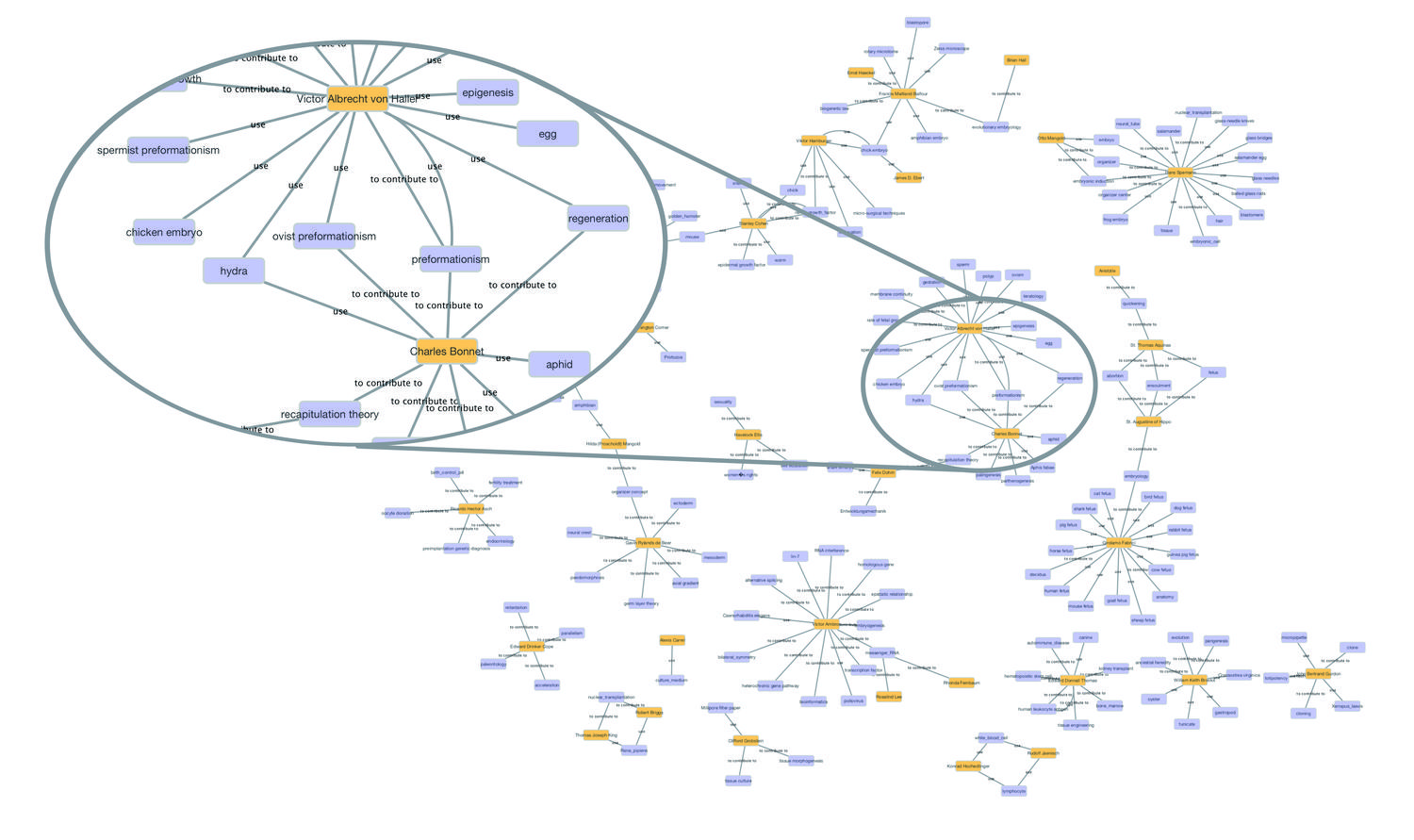
Fig. 21.2: Network of people and theories, organisms, and techniques created from 35 Embryo Project articles.
Statements made in such a context are considered to be true in the environment of the context. However, Macgregor and Ko do not define a structure for environments. In the case of Quadruples in the Quadriga System, the context is well-defined. It consists of three parts: the metadata of a resource (such as publication date or author), the annotation context (such as the creator of the annotations of a text), and the creation context (such as when annotations were uploaded to a shared repository).
The Quadriga System has several components that support the creation of Quadruple networks and their distribution with the following workflow. A researcher annotates each text of interest with a graph that represents their interpretation of that text. Such a graph consists of relationships between concepts that the researcher created according to the relevant statements of a text. Relevant information is in this context the information that the researcher classifies as being relevant. Next, additional information such as metadata of the text is attached to the graph. The researcher then uploads his graphs to a common repository. This repository holds graphs from several researchers working on possibly different projects. Once his graphs are uploaded, the Quadriga System enables the researcher to analyze them, incorporating or excluding specific graphs created by other researchers and projects.
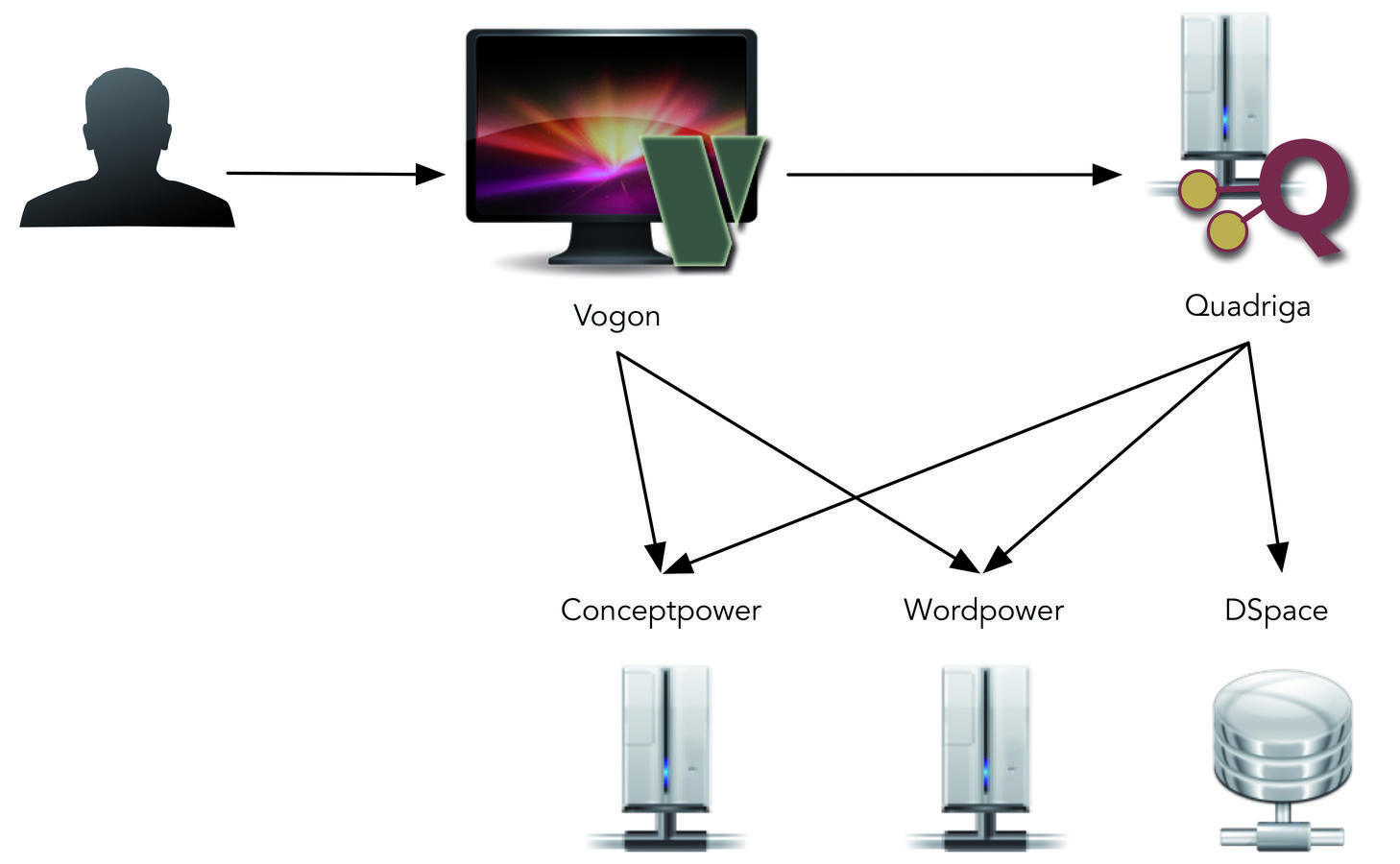
The Quadriga System consists of several independent components that interact with each other (see fig. 21.4). Each component has specific responsibilities. A user might directly interact with all components or with only a few depending on his role in a project. The component that users will likely interact with the most is Vogon. Vogon is a desktop application that enables users to annotate texts. This can be done with a text-based editor, in which users highlight the terms that they want to annotate, or using a graphical editor that lets users build a graph diagrammatically and then connect each node in the graph to the text.
Several annotations together form graphs. When a user has finished annotating a text, those graphs can be submitted to Quadriga, a network repository. Quadriga is the central component of the Quadriga System. It is a web application that provides functionality to review, annotate, store, and publish graphs consisting of Quadruples.
A basic element in the Quadriga System are texts: networks are created for texts, annotations link to positions in texts, and Quadriga manages graphs by associating them with specific texts. To use the Quadriga System to its full potential, documents that are being annotated using Vogon should be available to the whole system. This would allow, for example, visualization websites of annotation graphs to display the part of a text for which an annotation was created. In the Quadriga System, texts are therefore made available through a DSpace repository.4
The last two components in the Quadriga System are an online authority file service called Conceptpower and an online dictionary service called Wordpower. Both services are web applications. Quadriga as well as Vogon interact with these services through a web API (Application Programming Interface).5 In contrast, human users interact with Conceptpower and Wordpower through a website using a web browser.
Conceptpower is the authority file system used in the Quadriga System. Each entry in Conceptpower represents a concept and is identified by a URI. Given such a URI, an application can retrieve a concept’s properties, such as its type or the contents of the equals field. If a concept is missing in Conceptpower, a user can create a new entry in Conceptpower for the missing concept. Wordpower has many similarities with Conceptpower. As in Conceptpower, every entry in Wordpower is identified by a URI and, given the URI, other software applications can request information about a Wordpower entry. The biggest difference between the two services is that in Conceptpower each entry represents a specific meaning of a term. Even if two terms are the same, if they have different meanings there will be different entries in Conceptpower. In contrast, in Wordpower there is only one entry for a term and that entry specifies the normalized or correct spelling of a word.6
A typical annotation process using the Quadriga System looks like the following. A researcher starts by adding all texts he wants to annotate to Vogon. The user creates annotations for the texts and relates them to each other. During that step he queries Conceptpower and Wordpower for the terms and concepts he uses in his annotations. He also creates new entries in these two services if terms or concepts are missing. Once the researcher has finished annotating a text, he submits the annotation graphs to Quadriga for validation, publication, and visualization.
Conclusion
Quadruples, which are the underlying data structure of the Quadriga System, are contextualized triples of the form <subject - predicate - object - context>. Quadruples, in contrast to triples, store contextual information about a subject, predicate, object statement. Such contextual information contain, for instance, what text was annotated or who annotated a text. With this kind of data, it is possible to “zoom in and out” from the macro-level to the micro-level to allow scalable reading.
The Quadriga System operates on the meso-level between distance and close reading. Texts are annotated through close reading and examination of terms. However, all annotations are stored in a common repository, creating a large-scale dataset of annotation data (networks of Quadruples), which is available to other scholars. This dataset facilitates distant reading, which could assist in finding patterns and trends in the annotated corpus. However, distant reading in the context of the Quadriga System is limited by the number of annotated texts and the annotations created for those texts. Also, the annotation process itself is time-consuming and is likely to be restricted to a few texts of interest. It therefore might be practicable to use other distant reading techniques such as topic modeling on large text corpora to identify sub-corpora of interest for ingestion into the Quadriga System.
Compared to many large-scale text analysis methods such as topic modeling or co-citation analysis, the Quadriga System has the advantage of not only connecting concepts and texts but also qualifying that link. For instance, a co-citation analysis might suggest that two papers are related because they are co-cited often but it does not make any assertions about the kind of relationship between those papers. Do both papers make similar statements or does one reject the statements of the other? Similarly, topic modeling might connect two terms by placing them in the same topic. However, it does not specify what kind of relation exists between the two terms. Do texts that belong to a specific topic describe a similar relationship between these two terms, or are they using the same terms but contradict each other? The Quadriga System can answer such questions by qualifying the relationship between concepts. Two concepts are not only in relation to each other but are connected by a specific relationship. A scholar could use this property of the system by, for instance, identifying several papers connected to each other by co-citation analysis and then annotating these papers with Quadruples to determine their specific relationships. In contrast to traditional close reading methods of the identified papers, the Quadriga System would provide a researcher not only with a structured way of extracting relevant information that could then be computationally analyzed using, for instance, network analysis measures. It would also allow a researcher to publish the extracted information (the Quadruple networks) so that other scholars can examine it or use it for their own research.
The last point, publishing annotations or data to be shared among scholars, connects these tools to the vision that guided Peter Damerow throughout his distinguished career: openness and sharing of information. It also allows the history of science or any other field that uses such tools to benefit from an economy of scale that, in the fashion of big data, facilitates novel and surprising discoveries. Following again in Peter’s footsteps, who devoted his whole career to collaborations, we have built this system in order to enable different researchers to share their data, collaborate on interpretations, and to expose their work beyond the narrow disciplinary boundaries of a specific discipline. It has been our own experience that all really interesting and important problems require a multi-disciplinary approach, something that hopefully just got a bit easier because of tools such as the one presented here.
References
Baliga, Nitin S., Tray Ideker, Andrew Markiel, Owen Ozier, Paul Shannon, Jonathan T. Wang, Daniel Ramage, Nada Amin, and Benno Schwikovski (2003). Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Research 13(11):2498–2504.
Benson, David A., Ilene Karsch-Mizrachi, D. J. Lipman, Jim Ostell, and E. W. Sayers (2010). GenBank. Nucleic Acids Research 38. (Database issue):D46–51.
Jacobson, Daniel, Dan Woods, and Graig Brail (2011). The API Opportunity. APIs: A Strategy Guide:1–9.
Laubichler, Manfred and Jane Maienschein (2009). The Embryo Project: An Integrated Approach to History, Practices, and Social Contexts of Embryo Research. Journal of the History of Biology 43(1):1–16.
Lunenfeld, Peter, Anne Burdick, Johanna Drucker, Todd Presner, and Jeffrey Schnapp (2012). Distant/Close, Macro/Micro, Surface/Depth. In: Digital_Humanities. Cambridge: MIT Press, 39–40.
MacGregor, Robert and In-Young Ko (2003). Representing Contextualized Data using Semantic Web Tools. In: International Workshop on Practical and Scalable Semantic Systems PSSS1.
Mazza, Riccardo (2009). Introduction to Visual Representations. In: Introduction to Information Visualization. London: Springer.
Moretti, Franco (2009). Conjectures on World Literature. New Left Review 1:54–68.
Müller, Martin (2012). Scalable Reading. url: https://scalablereading.northwestern.edu/.
Peirson, Erick, Julia Damerow, and Manfred Laubichler (forthcoming). Don’t Panic! A Research System for Network-based Digital History & Philosophy of Science. In: Power of Networks: Prospects for Historical Network Research. Ed. by Florian Kerschbaumer, Linda v. Keyserlingk, Martin Stark, and Marten Düring. London: Routledge.
Footnotes
A term coined by Franco Moretti (2009). At that point Moretti used the term “distant reading” in the context of world literature and did not focus on computational methods to automatically extract information. However, the basic idea is the same: “[d]istant reading [...] allows you to focus on units that are much smaller or much larger than the text” (Moretti 2009, 57).
See http://www.cytoscape.org/ and Shannon (2003).
An API is “a way for two computer applications to talk to each other over a network (predominantly the Internet) using a common language that they both understand” (Jacobson, Woods, and Brail 2011, 5).
The normalization of a term could be singular for plural nouns, present tense for verbs, or simply the correct spelling of a word.